


Next: Indexing a Darwin Sequence
Up: Accessing a Darwin Sequence
Previous: The String Function
It is easy to search for all entries in a sequence database that
contain a specific pattern. The function
SearchDb(
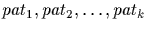 : { string, set(string) })
: { string, set(string) })
accepts a sequence of patterns where each pattern is either a
string or a set of string objects. When multiple
parameters are contained in a set, the SearchDb returns
the logical OR of the results, that is, it returns all Entry that contain any one of the patterns contained in the set. The comma symbol represents the logical AND of the
arguments. In this case, SearchDb returns only those Entry that contain all such patterns. SearchDb searches the
database currently assigned to the system variable DB.
> DB := ReadDb('Sample/SH2'): # load the SH2 database
# & assign it to DB
> SearchDb('mouse'); # returns all Entry() which contain
# the pattern 'mouse'.
Entry(7,8,10,13,17,22,26,32,34,37,39,47,64,67,69,73,78)
> SearchDb('SH3');
Entry(1,2,3,4,5,7,8,9,10,11,12,13,14,16,17,18,19,21,22,
23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,
42,43,44,45,46,47,48,49,50,51,52,53,54,55,56,57,58,59,60,
61,62,66,67,68,69,70,71,72,73,74,75,76)
> SearchDb('mouse', 'SH3'); # return all Entry() which contain
# both 'mouse' and 'SH3'.
Entry(7,8,10,13,17,22,26,32,34,37,39,47,67,69,73)
> SearchDb({'mouse', 'human'}); # return all Entry() which contain
> # human or mouse
Entry(2,3,7,8,9,10,12,13,16,17,21,22,23,25,26,30,31,32,
33,34,36,37,38,39,41,47,53,63,64,66,67,68,69,72,73,77,78)
> SearchDb('SH3', {'mouse', 'human'}); # return all Entry() containing
> # (SH3 and (mouse or human))
Entry(2,3,7,8,9,10,12,13,16,17,21,22,23,25,26,30,31,32,
33,34,36,37,38,39,41,47,53,66,67,68,69,72,73)
The SearchDb function is case insensitive, thus searching for
pattern 'mouse' is equivalent to searching for 'MOUSE'.
Any pattern is allowed as a parameter to the SearchDb function.
We can even search for peptide sequences.
> SearchDb('NGGYYI'); # a segment of the SH2 domain
Entry(8,24,25,26,27,35,38,39,40,43,70,71,72,73,74,75)
Note, however, that this is an exact match - in computer science
terms - a common substring search.
Gaston Gonnet
1998-09-15