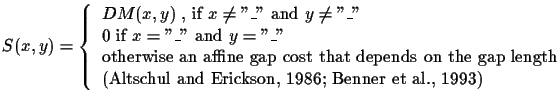
DM is an entry in a Dayhoff matrix [Schwarz and Dayhoff, 1979,Gonnet et al., 1992].
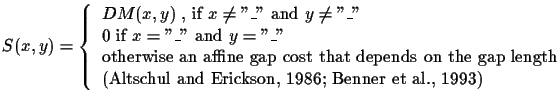
MPA scores must be equal to or less than the scores obtained from the OPA pairwise alignments. The CS method can be applied now again without the need for an explicitly defined evolutionary tree by repeating the procedure outlined above, but using MPA scores instead of OPA scores. The CS score of the MSA is the sum of MPA scores in the circular order C divided by two:
Note that if there is a deletion in both sequences, there is no penalty (score is zero) because that deletion happened in some ancestor, and its penalty has been counted already 2.
|