Definition 1.5
The score of the induced MSA-derived pairwise alignment
MPA(ai, aj) of two sequences
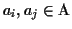
is the
score of the alignment of the two strings
ai and
aj, where
any scoring function for pairwise alignments can be used. Opposing
gaps may be removed.
 where
Cn+1 = C1, and C is a circular order
with respect to the evolutionary tree of
where
Cn+1 = C1, and C is a circular order
with respect to the evolutionary tree of